From Generalist to Specialist: Fine-Tuning GPT-2 for Biomedical Instruction Following
In the rapidly evolving world of Large Language Models (LLMs), a common challenge persists: general-purpose models often struggle with domain-specific tasks. While they can chat about anything from history to cooking, they frequently "hallucinate" or lack depth when asked about niche fields like medicine or law.
In this project, I embarked on an experiment to transform a standard GPT-2 Small model into a biomedical expert capable of following specific instructions. Using a staged transfer learning pipeline, I took the model from a confused generalist to a domain-aware assistant.
Here is the step-by-step journey of how I built this biomedical specialist.
The Objective
The goal was to demonstrate staged transfer learning. Instead of training a model from scratch (which requires massive compute), I fine-tuned a pre-trained model through distinct phases to inject domain knowledge (biomedical) and behavioral patterns (instruction following).
Phase 1: Establishing a Baseline
Before training, I needed to quantify how "ignorant" the base GPT-2 model was regarding biomedical texts. I evaluated the pre-trained weights on a domain-specific corpus derived from PubMed abstracts.
Metric: Perplexity (PPL) — a measure of how "surprised" the model is by new text.
Result: The baseline model achieved a Perplexity of 31.86.
Qualitative Analysis: When prompted with technical sentences, the model produced fluent but scientifically nonsensical text. It lacked the deep understanding required for the field.
Phase 2: Domain Adaptation
To turn the model into a "specialist," I performed Causal Language Modeling (CLM) on the biomedical corpus. This phase essentially sent the model to "medical school." I used the Hugging Face Trainer API with mixed-precision training (fp16) to optimize GPU memory usage.
# Training arguments for Domain Adaptation
training_args = TrainingArguments(
output_dir="./gpt2trained",
num_train_epochs=3,
per_device_train_batch_size=4,
gradient_accumulation_steps=4, # Simulating larger batch sizes
fp16=True, # Mixed precision for GPU memory efficiency
logging_steps=50,
save_total_limit=2
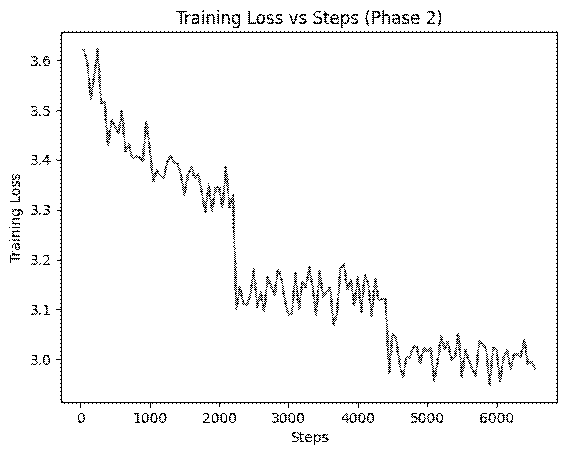
)Results: The training loss curve showed a steady decrease, indicating successful learning of domain patterns. Most importantly, the Perplexity dropped significantly to 16.80 (down from 31.86), proving the model was now much more comfortable with medical terminology.
Phase 3: The Discriminator
A robust AI system needs to distinguish between relevant and irrelevant information. In Phase 3, I repurposed the domain-adapted model as a binary classifier.
To train this, I needed a balanced dataset. I used biomedical abstracts as positive samples and generated synthetic negative samples (general daily life sentences) to force the model to learn the difference.
# Generating random general-domain sentences
# to contrast with biomedical abstracts
neg_set = set()
while len(neg_set) < 250:
s = f"{random.choice(subjects)} {random.choice(verbs)} to {random.choice(places)}..."
neg_set.add(s)Result: After fine-tuning the classification head, the model achieved 100% accuracy on the test set. This confirmed that the model had learned distinct linguistic features separating biomedical text from general content.
Phase 4: Instruction Fine-Tuning
The final and most critical phase was teaching the model to act as an assistant. Mere knowledge isn't enough; the model needed to understand how to respond to questions. I curated a dataset of 50 instruction-response pairs and formatted them into a structured prompt. This format explicitly teaches the model where the user's query ends and where its response should begin.
def format_example(ex):
"""
Formats the input to train the model on instruction following.
"""
return (
"### Instruction:\n"
+ ex["instruction"].strip()
+ "\n### Response:\n"
+ ex["response"].strip()
+ tokenizer.eos_token
)Final Results: The Transformation
The difference between the base model and the final instruction-tuned model is night and day.
Prompt: "We performed Cox proportional hazards regression..."
Model Output: "...to estimate the hazard ratio (HR) for selected subgroups... The HR for the lowest group is 3.08..."
Analysis: The model hallucinates statistics and continues the sentence blindly.
2. After Instruction Tuning (Phase 4)
Prompt: "What is Cox proportional hazards regression?"
Model Output: "Cox proportional hazard regression measures the influence of a given outcome on the likelihood that it will occur..."
Analysis: The model now acts as an intelligent assistant, providing a correct and relevant definition.
Conclusion
This project demonstrates the power of Transfer Learning and Instruction Tuning. By taking a small, general-purpose model (GPT-2) and guiding it through specific training phases, we can build specialized tools for complex domains like healthcare, all with limited computational resources.